TRANSLATIONAL GENOMICS
Translational genomics is a new emerging science combining translational and clinical research with statistics, genomics, clinical informatics, bioinformatics, medical informatics, information technology, and mathematics together. One of our current research goals is to apply/develop appropriate computational tool to analyze the high-throughput omics data. We are currently working with Dr. Qi Chen's Lab to identify the “disease RNA code” composed of non-canonical small non-coding RNAs (e.g. tRNA-derived small RNAs [tsRNAs], rRNA-derived small RNAs [rsRNAs], and YRNA-derived small RNAs [ysRNAs]), which potentially paves a new avenue for future molecular diagnosis/prognosis in precision medicine.
NON-CODING RNA BIOINFIRMATICS
With the development of high-throughput sequencing technology, it has been revealed that, in eukaryotes, only a small proportion of transcripts carry protein-coding potential, while the majority of the genome is transcribed to produce non-coding RNAs. Non-coding RNAs are often involved in regulating gene expression at the transcriptional and/or post-transcriptional levels. It is my research interest to investigate the non-canonical small non-coding RNAs, such as tsRNAs, rsRNAs, ysRNAs, and so on, which are markedly under investigated but miraculously involved in intergenerational/transgenerational epigenetic inheritance.
NON-NEUTRAL EVOLUTION AT SYNONYMOUS SITES
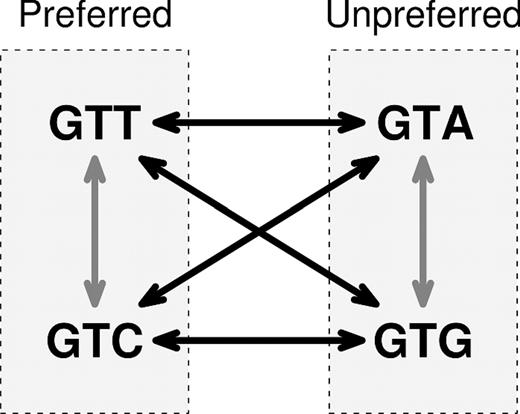
Synonymous mutations (so called silent mutations) are the nucleotide change in a codon of a gene, such that the produced protein sequence is not modified. When a synonymous or silent mutation occurs, the change is often assumed to be neutral and thus have no effect on fitness. However, increasing evidence indicates that synonymous mutations have significant consequences for cellular processes in all taxa. Evolutionary research on synonymous sites is becoming increasingly important as these analyses make their way into clinical utilizations of academic results, which has numerous implications not only for the understanding of basic biology but also for the diagnosis and treatment of genetic diseases.